MFront implementation¶
To give a first introduction to writing material behaviors using MFront, we will implement a very simple linear elastic model :
where and are the Lamé coefficients.
The parameters of the model will be Young’s modulus and Poisson’s ration , while the output will be the Cauchy stress tensor .
Of course, we could use one of Mfront’s standard brick for this purpose, but the idea here is to define everything ourselves.
Here is a look at the corresponding elasticity.mfront file:
@DSL DefaultGenericBehaviour;
@Author Maxime Pierre;
@Date 22/11/2025;
@Behaviour Elasticity;First, we specify the domain specific language (DSL) that we want to use.
It will condition what keywords we will have access to in the rest of the file.
After optionally declaring an author and a date, we give a name to our behaviour (Elasticity).
Next, we define the input and output variables of our model : the strain tensor (a gradient of the displacement), and its associated flux which is the stress tensor .
@Gradient StrainStensor εᵗᵒ;
εᵗᵒ.setGlossaryName("Strain");
@Flux StressStensor σ;
σ.setGlossaryName("Stress");Then, we define the material properties which must be passed to the behaviour:
@MaterialProperty stress E;
E.setGlossaryName("YoungModulus");
@MaterialProperty real ν;
ν.setGlossaryName("PoissonRatio");Naming variables in MFront
MFrontNote that all the names of variables and properties used so far are very common, and part of the standard glossary of MFront. They are therefore applied using the setGlossaryName method. For names which are not part of the glossary, any name can be applied using the setEntryName method instead.
Now, we define the blocks composing the tangent operator. In our simple case, their is only one, , which corresponds to the stiffness tensor .
@TangentOperatorBlocks{∂σ∕∂Δεᵗᵒ};
@ProvidesTangentOperator ;Finally, we enter the core of the behaviour with the Integrator block.
This is where the behaviour of the material is actually calculated.
MFront provides some helper functions to calculate Lamé’s coefficients from and .
Then, we calculate the stress with an incremental formulation (note that in the case of elasticity, a total formulation would also have been acceptable).
The last part gives the one tangent operator block that was declared above.
@Integrator{
const auto λ = computeLambda(E, ν);
const auto μ = computeMu(E, ν);
σ += λ ⋅ trace(Δεᵗᵒ) ⋅ I₂ + 2 ⋅ μ ⋅ Δεᵗᵒ ;
if (computeTangentOperator_) {
∂σ∕∂Δεᵗᵒ = λ ⋅ (I₂ ⊗ I₂) + 2 ⋅ μ ⋅ I₄;
}
}Compiling a behaviour¶
Once the .mfront file is written, we need to compile it to create a shared library that can then be used in other contexts (mtest, python, FEniCS...).
This can be done from the terminal using the following command:
mfront --obuild --interface=generic elasticity.mfrontHere, we used the generic interface, which is the one used to plug the constitutive laws into python or FEniCS using MGIS, the MFront Generic Interface Support.
The command creates or updates a shared library src/libBehaviour.so on Linux or src/libBehaviour.dylib on MacOS (note the difference in extension for dynamic libraries).
Testing the behaviour with mtest¶
@Behaviour<Generic> 'src/libBehaviour.dylib' 'Elasticity';
@MaterialProperty<constant> 'YoungModulus' 1e+10 ;
@MaterialProperty<constant> 'PoissonRatio' 0.2 ;@ExternalStateVariable 'Temperature' 293.15 ;
@ImposedDrivingVariable 'StrainXX' 0.0;
@ImposedDrivingVariable 'StrainYY' 0.0;
@ImposedThermodynamicForce 'StressZZ' {0.0: 0.0, 1.0: -1e7};
@Times {0.0, 1.0 in 10};mtest elasticity.mtestInteracting with the behaviour in python¶
Using the mtest package in python, it is possible to interact with a compiled MFront behaviour.
First, we have to set the behaviour for our test as well as the different material properties that are necessary. We chose Young’s Modulus as 10 MPa and Poisson’s ration as 0.2.
Source
from sys import platform
import std
import tfel.math
import numpy as np
from mtest import (
MTest,
VerboseLevel,
PredictionPolicy,
setVerboseMode,
MTestCurrentState,
MTestWorkSpace,
Behaviour,
)
def lib_ext() -> str | None:
if platform == "linux":
return ".so"
elif platform == "darwin":
return ".dylib"
setVerboseMode(VerboseLevel.VERBOSE_LEVEL2)test = MTest()
state = MTestCurrentState()
workspace = MTestWorkSpace()
# Setting the behaviour
behaviour = Behaviour("generic", f"../MFront_library/src/libBehaviour{lib_ext()}", "Elasticity","Tridimensional")
test.setBehaviour("generic", f"../MFront_library/src/libBehaviour{lib_ext()}", "Elasticity")
# Setting the material properties
test.setMaterialProperty("YoungModulus", 1e10)
test.setMaterialProperty("PoissonRatio", 0.2)Next, we set the conditions for the test. The setImposedGradient method allows to control the different components of the gradients (here strain) by their name, and the setImposedThermodynamicForce method does the same for thermodynamic forces (here stress). A single value can be provided, or a dictionary with several entries to have a variation over time, similar to the mtest syntax.
We impose oedometric conditions on the sample: and deformation are prevented while we apply an axial load up the 10 MPa in compression.
test.setExternalStateVariable("Temperature", 293.15)
test.setImposedGradient("StrainXX", 0.0)
test.setImposedGradient("StrainYY", 0.0)
test.setImposedThermodynamicForce("StressZZ", {0.0: 0.0, 1.0: -1e7})
test.completeInitialisation()
test.initializeCurrentState(state)
test.initializeWorkSpace(workspace)We perform a simulation with 10 steps between times 0 and 1, and store the results to show the stress-strain curve of the oedometric test. Obviously, this is a very simple test and the results is a simple line with slope .
t = np.linspace(0,1,10)
epszz_index = behaviour.getGradientComponentPosition("StrainZZ")
sigzz_index = behaviour.getThermodynamicForceComponentPosition("StressZZ")
epszz = [0.0]
sigzz = [0.0]
for i in range(len(t)-1):
test.execute(state, workspace, t[i], t[i+1])
epszz.append(-state.e1[epszz_index])
sigzz.append(-state.s1[sigzz_index])Source
import matplotlib.pyplot as plt
plt.plot(epszz, sigzz)
plt.xlabel(r"$\varepsilon_{zz}$ [Pa]")
plt.ylabel(r"$\sigma_{zz}$ [-]")
plt.show()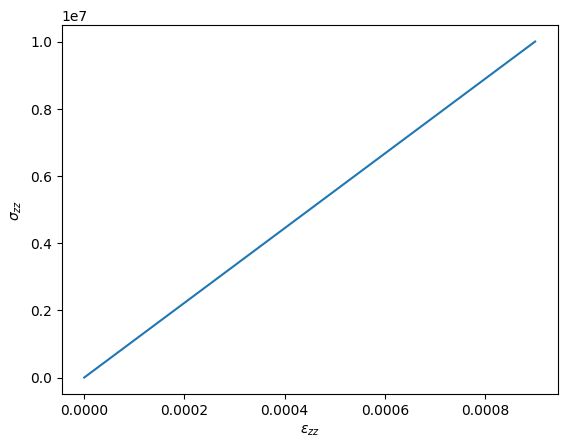
We can compare the resulting oedometric modulus from the simulation with the analytical formula:
print(f"Modulus from simulation: {sigzz[-1]/epszz[-1]/1e9:.2f} GPa")
print(f"Analytical oedometric modulus: {1e10*(1-0.2)/(1+0.2)/(1-2*0.2)/1e9:.2f} GPa")Modulus from simulation: 11.11 GPa
Analytical oedometric modulus: 11.11 GPa